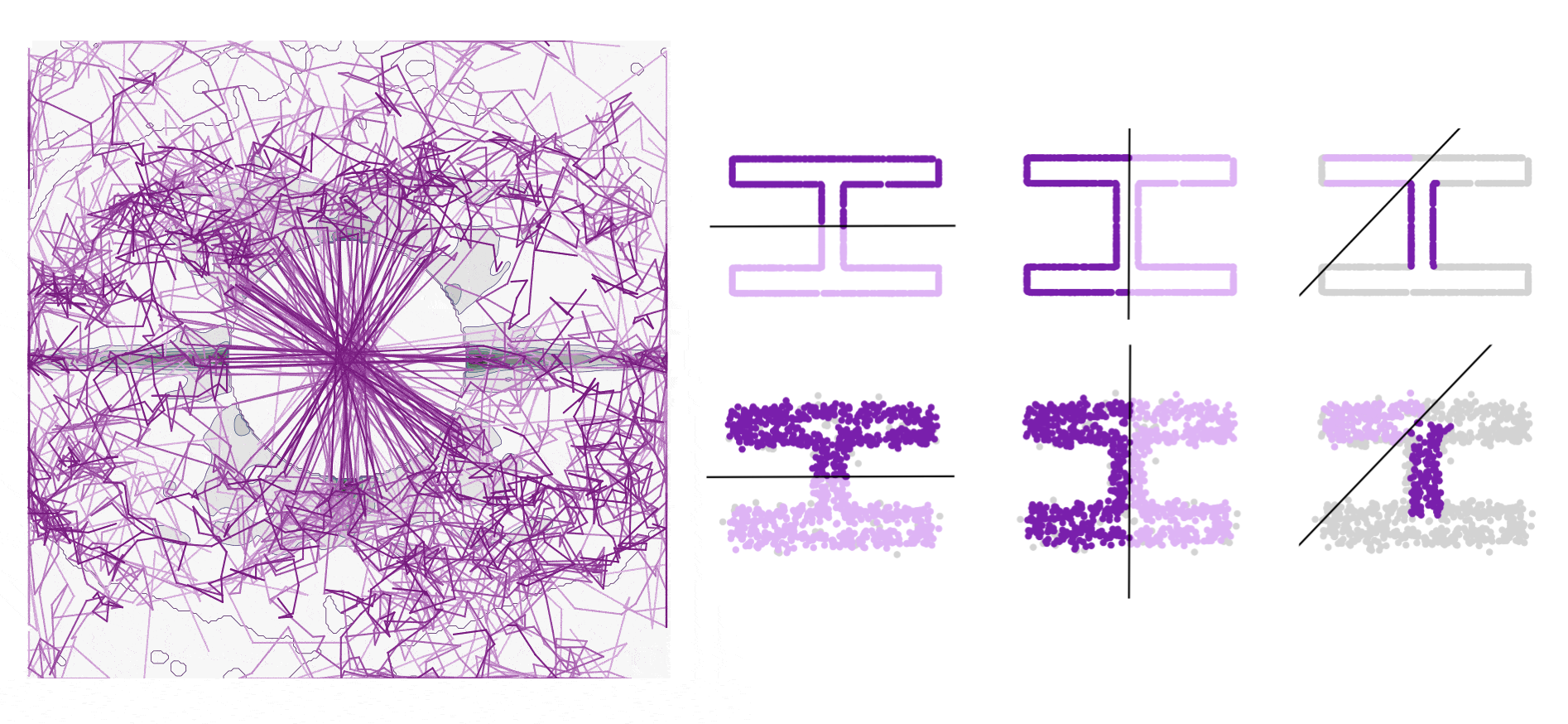
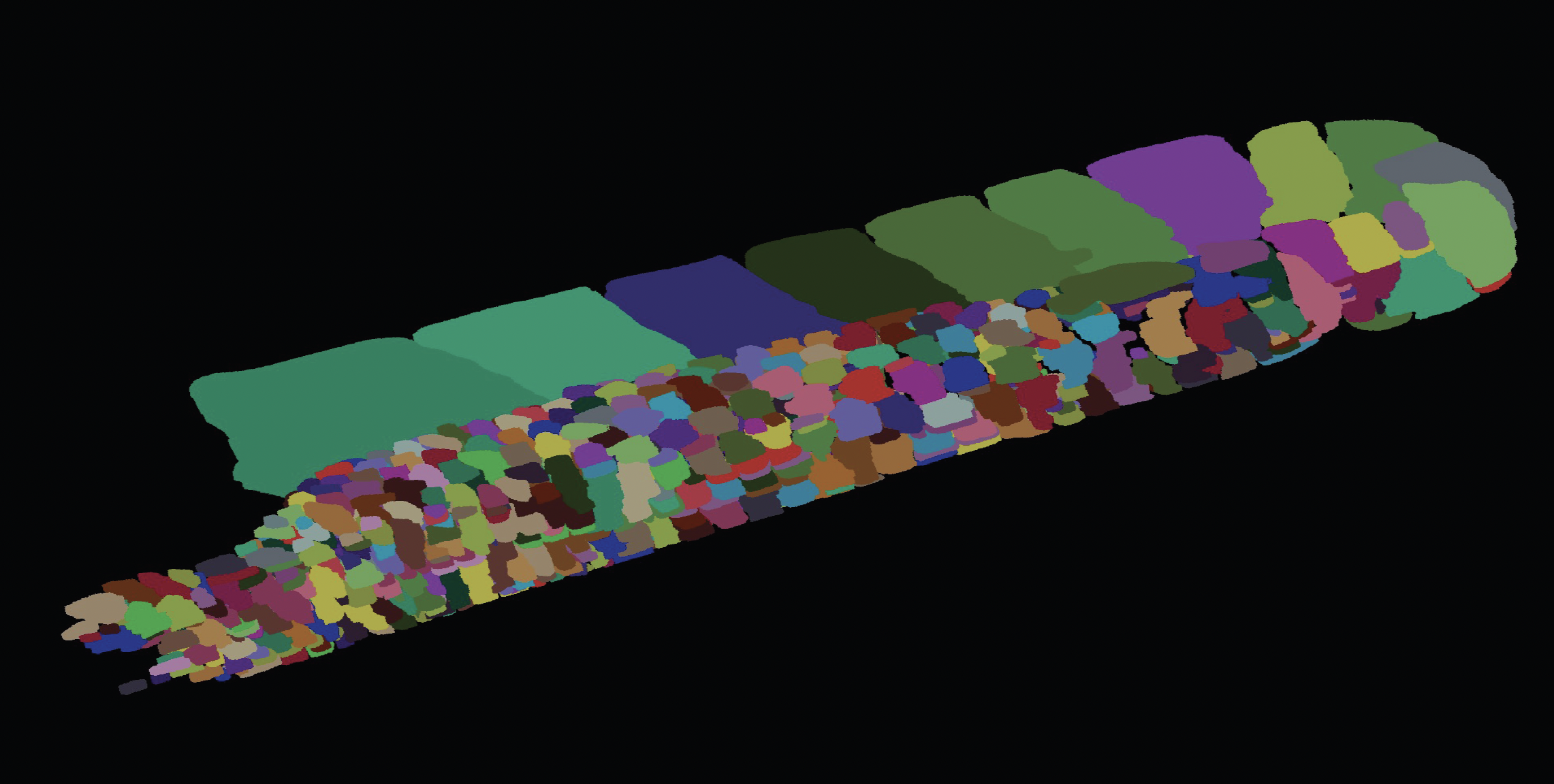
Robust Symmetry Detection via Riemannian Langevin Dynamics
Jihyeon Je*, Jiayi Liu*, Guandao Yang*, Boyang Deng*, Shengqu Cai, Gordon Wetzstein, Or Litany, Leonidas Guibas
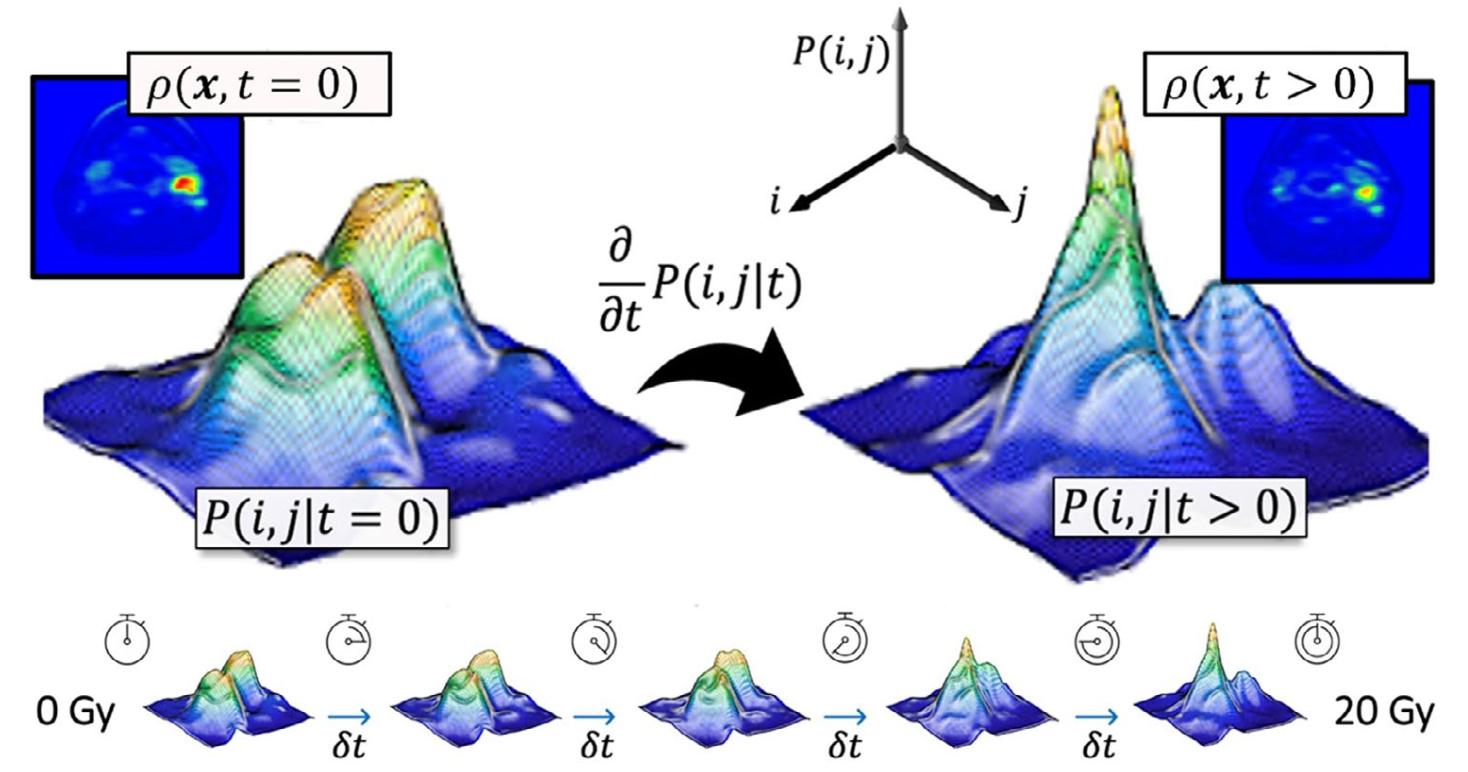
Connecting Langevin dynamics with Meanshift for more robust symmetry detection on 2D and 3D shapes